Originally posted by dannydesloover
View Post
Seqanswers Leaderboard Ad
Collapse
Announcement
Collapse
No announcement yet.
X
-
I am also interested in using the QIAxcell for this purpose, could you share some expertize with us?Originally posted by garciad1 View PostWe recently installed a Qiaxcel in our lab, but the training was horrible. Could you tell me what run parameters, marker, etc. you use for running NGS libraries for sizing etc.? I can't seem to find any references for using the system for this purpose. --Thanks!
Comment
-
Can Qiaxcel be used instead of Bioanalyzer for Library QC?
I'm hoping to purchase a MiSEQ in the next few months and wondering whether I need a Bioanalzyer as I already have a Qiaxcel - can you use the Qiaxcel for library QC? Do you have any SOPs for doing this?Originally posted by adaptivegenome View PostI use the Qiaxcel system. It works for me as well as the Bioanalyzer but the sample cost is about 4 times lower. It does use gel capillaries and the array lasts for only 6 months at a time so if you are not a high volume user, it might not be as cost effective as it is for me.
Thanks,
k
Comment
-
I believe the Qiaxcell injects markers independently of sample, so it will deliver size data, but you will need a fluorometer (Qubit) to get accurate concentration measurements. Ionic strength differences between samples and standards impact injection efficiency. Normalization is required to measure concentration.
The Fragment Analyzer offers qualitative kits that inject markers independently of sample like the Qiaxcel. These kits are inexpensive and useful for sizing applications like PCR screening, genotyping, RFLP, microsatellites, etc. They are also useful for relative concentration measurements of peaks within a sample. IDT just published a CRISPER/Cas9 cleavage assay for their gene block products using these kits on the Fragment Analyzer:
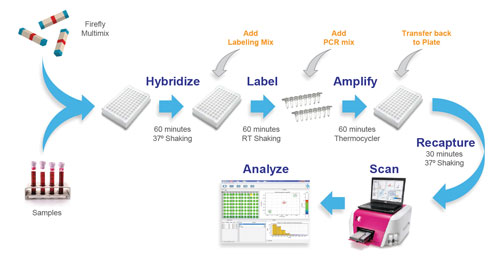 To lessen CRISPR’s resistance to high-throughput application, Integrated DNA Technologies introduces fully synthesized sgRNA transcription fragments.
To lessen CRISPR’s resistance to high-throughput application, Integrated DNA Technologies introduces fully synthesized sgRNA transcription fragments.
The Fragment Analyzer also has quantitative kits that mix markers with samples like the BioAnalyzer. This enables use of the marker molecules as a normalization factor with reference to the molecules in the ladder as concentration standards. The system is easy to use and reliable with automated gel cycling and parallel analysis (not rapid serial injection). The different Fragment Analyzer quantitative kits work for small Illumina type NGS libraries as well as larger PacBio and nanopore type libraries and genomic DNA that exceed the size range of the BioAnalyzer. Resolution and sensitivity are excellent. Its a solid analytical platform for all kinds of DNA and RNA applications.
Comment
Latest Articles
Collapse
-
by seqadmin
The sequencing world is rapidly changing due to declining costs, enhanced accuracies, and the advent of newer, cutting-edge instruments. Equally important to these developments are improvements in sequencing analysis, a process that converts vast amounts of raw data into a comprehensible and meaningful form. This complex task requires expertise and the right analysis tools. In this article, we highlight the progress and innovation in sequencing analysis by reviewing several of the...-
Channel: Articles
Yesterday, 07:48 AM -
-
by seqadmin
The field of epigenetics has traditionally concentrated more on DNA and how changes like methylation and phosphorylation of histones impact gene expression and regulation. However, our increased understanding of RNA modifications and their importance in cellular processes has led to a rise in epitranscriptomics research. “Epitranscriptomics brings together the concepts of epigenetics and gene expression,” explained Adrien Leger, PhD, Principal Research Scientist...-
Channel: Articles
04-22-2024, 07:01 AM -
ad_right_rmr
Collapse
News
Collapse
| Topics | Statistics | Last Post | ||
|---|---|---|---|---|
|
Started by seqadmin, Today, 06:57 AM
|
0 responses
9 views
0 likes
|
Last Post
by seqadmin
Today, 06:57 AM
|
||
|
Started by seqadmin, Yesterday, 07:17 AM
|
0 responses
14 views
0 likes
|
Last Post
by seqadmin
Yesterday, 07:17 AM
|
||
|
Started by seqadmin, 05-02-2024, 08:06 AM
|
0 responses
19 views
0 likes
|
Last Post
by seqadmin
05-02-2024, 08:06 AM
|
||
|
Started by seqadmin, 04-30-2024, 12:17 PM
|
0 responses
23 views
0 likes
|
Last Post
by seqadmin
04-30-2024, 12:17 PM
|

Comment