ChIPBase v2.0 is designed for studying transcription factor binding sites and motifs, and decoding transcriptional regulatory networks of non-coding RNAs (lncRNAs, miRNAs, snoRNAs, tRNAs...) and protein-coding genes (PCGs) from ~10,200 ChIP-seq data (ChIP-seq, ChIP-exo, MNChIP-seq) in 10 speices (Human, Mouse, Fruitfly, Worm, A. Thaliana, Yeast, Rat, Zebrafish, X. tropicalis, Chicken).
We identified thousands of Binding Motif Matrices and their binding sites from ChIP-seq data of DNA-binding proteins and predicted millions of transcriptional regulatory relationships between transcription factors and genes (lncRNAs, microRNAs, protein-coding genes, snoRNAs, tRNAs).
We constructed Regulator module to predict hundreds of transcription factors that were involved in or affected transcription of ncRNAs and protein-coding genes.
Moreover, we built a web-based tool, Co-Expression, to explore the co-expression patterns between DNA-binding proteins and various types of genes by integrating the gene expression profiles of ~10,000 tumor samples and ~9,100 normal tissues or cell lines.
ChIPBase also provides a ChIP-Function tool and a genome browser to predict functions of diverse genes and visualize various ChIP-seq data. This study will greatly expand our understanding of the transcriptional regulations of ncRNAs and protein-coding genes.
ChIPBase paper is available at http://nar.oxfordjournals.org/conten...ar.gkw965.full
ChIPBase is freely available at http://rna.sysu.edu.cn/chipbase/.
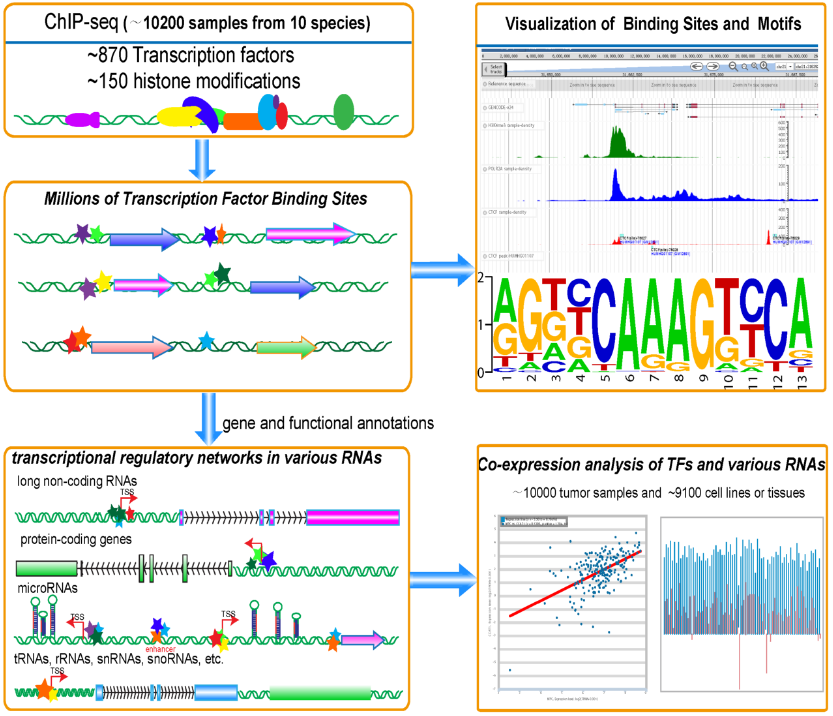
We identified thousands of Binding Motif Matrices and their binding sites from ChIP-seq data of DNA-binding proteins and predicted millions of transcriptional regulatory relationships between transcription factors and genes (lncRNAs, microRNAs, protein-coding genes, snoRNAs, tRNAs).
We constructed Regulator module to predict hundreds of transcription factors that were involved in or affected transcription of ncRNAs and protein-coding genes.
Moreover, we built a web-based tool, Co-Expression, to explore the co-expression patterns between DNA-binding proteins and various types of genes by integrating the gene expression profiles of ~10,000 tumor samples and ~9,100 normal tissues or cell lines.
ChIPBase also provides a ChIP-Function tool and a genome browser to predict functions of diverse genes and visualize various ChIP-seq data. This study will greatly expand our understanding of the transcriptional regulations of ncRNAs and protein-coding genes.
ChIPBase paper is available at http://nar.oxfordjournals.org/conten...ar.gkw965.full
ChIPBase is freely available at http://rna.sysu.edu.cn/chipbase/.
